Cancer Microbiome Research
Microbial Communities Influence Host Immune Response
Microbes living in the host can influence cancer progression and treatment efficacy. Diet and drugs can disrupt microbiome diversity, and key species in the microbiome can cause local or systemic influences on host immunity.1,2
There is hope that future treatments may combine existing cancer therapies with methods to encourage growth of beneficial microbes or eliminate harmful ones. As NGS-based research continues to explore host-microbiome interactions, Illumina strives to evolve genomic technologies that complement and enable the promise of this field.
Cancer Immunotherapy and the Role of the Microbiome
The effectiveness of modern cancer immunotherapies can be influenced by gut microbiome composition.
Microbes Can Exert Varied Effects on Cancer Progression
Microbes that influence cytokine release and inflammation can exert either a positive or negative effect on tumor growth, or the ability of the immune system to suppress tumor growth.
NGS allows profiling of microbial communities in different contexts, which is critical for identifying species or conditions that may be targeted for developing new therapeutic approaches to cancer.
Metagenomics Enabled by NGS Applications
NGS methods have revolutionized the study of the microbiome.3 Without the need to culture or clone individual organisms, NGS enables simultaneous comparison of thousands of microbial community species in samples from healthy individuals vs. those with cancer. With the development of bioinformatic tools to manage large volumes of novel data, this shift from single organism analysis enables accurate assessment of species diversity and measurement of dynamic fluctuations in microbial communities.
The Microbiome Emerges as a Key Player in Cancer
An overview of how the microbiome influences cancer and immunotherapy, and the role NGS plays to help advance research in this expanding field.
NGS Methods to Study the Role of the Microbiome in Cancer
The ability of NGS to analyze complex microbial communities may help to uncover new mechanisms of host–microbe interactions that promote cancer or promote drug efficacy.
The throughput and cost savings of NGS have fueled metagenomics studies capable of surveying the genomes of entire microbial communities.
NGS methods that can provide critical genetic insight into the microbiome:
Shotgun Metagenomic Sequencing
A DNA sequencing method that enables comprehensive sampling of all genes in all organisms in complex microbiome samples.
16S rRNA Sequencing
A 16S ribosomal RNA gene sequencing method used to identify and compare bacteria present within a given sample.
Microbial Metatranscriptomics
Analysis of all RNAs encoded by a group of microorganisms within a complex sample.
How Microbiome Multiomics Can Bolster Human Health
Sequencing technologies are enabling a deeper analysis of the gut's microbiome. Researchers can now explore what our microbial inhabitants are doing and how they contribute to, or protect from, diseases such as cancer and autoimmune disorders.
Read Article
Cancer Microbiome Analysis Workflow
With high throughput and high sensitivity, NGS enables identification of thousands of microbial species in cancer vs. normal samples.
Click on the below to view products for each workflow step.
Prepare sequencing ready libraries of bacteria, viruses, and other microbes to analyze transcriptome and metatranscriptome information.
Illumina Stranded Total RNA Prep with Ribo-Zero PlusStreamlined, cost-efficient, and scalable solution for total RNA analysis.
A fast, integrated workflow for a wide range of applications, from human whole-genome sequencing to amplicons, plasmids, and microbial species.
Speed, accuracy and simplicity for far-reaching applications in microbiology.
NextSeq 550 SystemFlexible benchtop sequencer with speed and simplicity for everyday genomics.
These cost-efficient, user-friendly, mid-throughput benchtop sequencers offer extreme flexibility to support new and emerging applications.
NovaSeq 6000 SystemHigh throughput and low cost for production-scale genomics.
Performs taxonomic classification of 16S rRNA targeted amplicon reads using an Illumina-curated version of the GreenGenes taxonomic database.
DRAGEN Metagenomics PipelinePerforms taxonomic classification of reads and provides single sample and aggregate reporting.
Data management and simplified bioinformatics for labs getting started and for rapidly scaling up sequencing operations.
Gene-Level Metagenomics for Microbiome Research
Dr. Samuel Minot from the Fred Hutchinson Cancer Research Center discusses a novel gene-level bioinformatics approach for identifying microbial species and strains associated with human diseases such as cancer, from metagenomic sequencing datasets.
View Webinar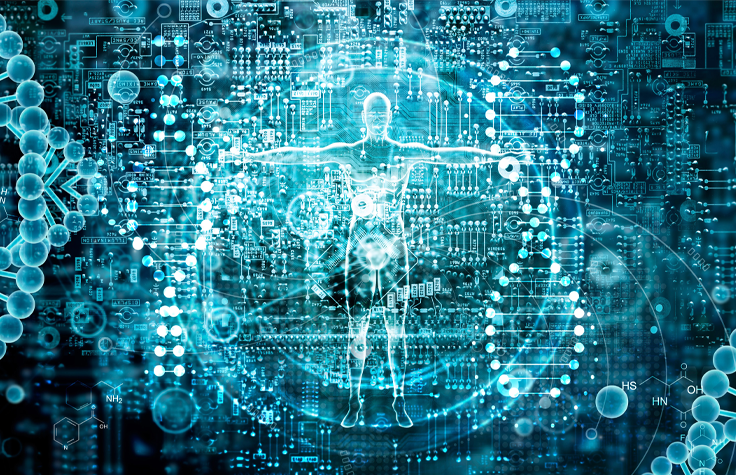
NGS is Revealing the Mysteries of the Microbiome
Researchers at Microba are investigating the genomes of microbes to improve our understanding of human health, disease, and microbial evolution.
Read Interview
Related Applications
Analyze Gene Expression and Chromatin Accessibility
Unify single-cell gene expression and ATAC-Seq to help reveal cellular mechanisms driving gene regulation.
Read Tech NoteMeasure Protein Expression Along with RNA-Seq
Learn how to incorporate protein detection into bulk RNA-Seq and develop a workflow for BEN-Seq, providing a holistic approach to cell analysis.
Read App NoteAdditional Resources

HPV and Skin Cancer
Illumina sequencers offer deep coverage to identify novel HPV types correlated with non-melanoma skin cancers.
Interested in receiving newsletters, case studies, and information on cancer genomics?
Sign UpReferences
- Zama D, Biagi E, Masetti R, et al. Gut microbiota and hematopoietic stem cell transplantation: where do we stand? Bone Marrow Transplant. 2017;52 (1):7-14.
- Schwabe R, Jobin C. The microbiome and cancer. Nat Rev Cancer. 2013;13 (11):800-812.
- Franzosa EA, Hsu T, Sirota-Madi A, et al. Sequencing and beyond: integrating molecular 'omics' for microbial community profiling. Nat Rev Microbiol. 2015;13(6):360-372.
