Target Enrichment
What is Target Enrichment?
Target enrichment works by capturing certain genes or genomic regions of interest by hybridization to target-specific biotinylated probes, which are then isolated by magnetic pulldown and sequenced with next-generation sequencing (NGS).
Illumina library prep and enrichment kits offer a complete solution for targeted enrichment. Illumina DNA Prep with Enrichment combines bead-linked transposome-mediated tagmentation chemistry with hybrid-capture target enrichment, reducing workflow time and producing an accurate, reliable assay for target enrichment resequencing for genomic DNA, tissue, blood, saliva and FFPE samples. Illumina Cell-Free DNA Prep with Enrichment offers fast, flexible library prep for highly sensitive mutation detection from cfDNA samples.

Benefits of NGS Target Enrichment
Hybridization-based enrichment is a useful strategy for analyzing specific genetic variants in a given sample.
Target enrichment allows researchers to reliably sequence exomes or large numbers of genes (> 50 genes) using robust and straightforward workflows. It delivers dependable results across a wide range of input types and quantities. Uses include:
- Human whole-exome sequencing (WES)
- Predesigned (fixed) sequencing panels for cancer, genetic disease and infectious disease research
- Custom enrichment panels for targeted sequencing

Target Enrichment vs. Other Resequencing Strategies
Hybrid capture-based target enrichment lets researchers target regions of the genome that are relevant to their specific research interests. Targeted resequencing delivers higher discovery power (ability to identify novel variants), and higher mutation resolution than PCR or Sanger sequencing.
Compared to PCR-based amplicon sequencing, hybridization-based enrichment sequencing can target a higher amount of total gene content and support more comprehensive profiling of all variant types. The larger amount of total gene content allows for the characterization of both known and novel variants for discovery-related applications.
Learn more about:
Benefits of Target Enrichment vs. Amplicon Sequencing*
Target Enrichment |
Amplicon Sequencing |
| Larger gene content, typically > 50 genes | Smaller gene content, typically < 50 genes |
| More comprehensive profiling for all variant types | Ideal for analyzing single nucleotide variants and insertions/deletions (indels) |
| More comprehensive method, but with longer hands-on time and turnaround time** | More affordable, easier workflow |
* Data on file, Illumina 2018.
** The turnaround time is for library prep assay time (DNA to finished library).
Recommended Target Enrichment Workflow
User-friendly automation-compatible workflows provide fast and flexible solutions for researchers. On-bead tagmentation chemistry enables support for a wide range of DNA input amounts, various sample sizes, and a broad range of applications.
Featured Infectious Disease Target Enrichment Solutions
Viral Surveillance Panel v2
Whole-genome sequencing and characterization of >200 viruses and subtypes including SARS-CoV-2, influenza, arboviruses, and hepatitis with target enrichment.
Respiratory Pathogen ID/AMR Panel
This NGS-based target enrichment workflow targets respiratory pathogens and antimicrobial resistance alleles, and offers simplified data analysis.
Empowering access for groundbreaking genomic discoveries
Illumina benchtop sequencing systems are making NGS technology more accessible to laboratories worldwide. Learn how these systems provide the speed, power, and flexibility to make breakthroughs in microbiology, cancer research, and more. The MiSeq i100 Series or NextSeq 1000 and NextSeq 2000 Systems can help make your NGS research goals within reach.
Download eBook
Featured Products
Related Solutions
Complex Disease Research
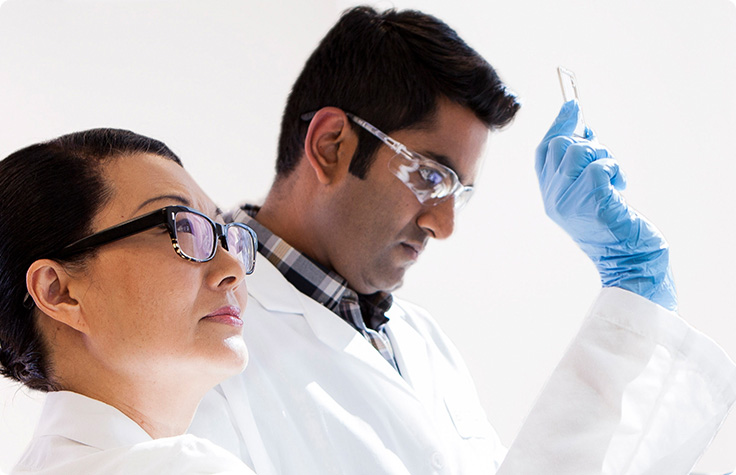

Array and NGS methods enable global genetic analyses to facilitate complex disease studies.
Learn MoreSomatic Variant Analysis
Perform somatic variant profiling studies of tumor specimens.
Learn MoreExome Sequencing
Investigate the protein-coding regions of the genome to uncover genetic influences on disease and population health.
Learn MoreInfectious Disease Applications
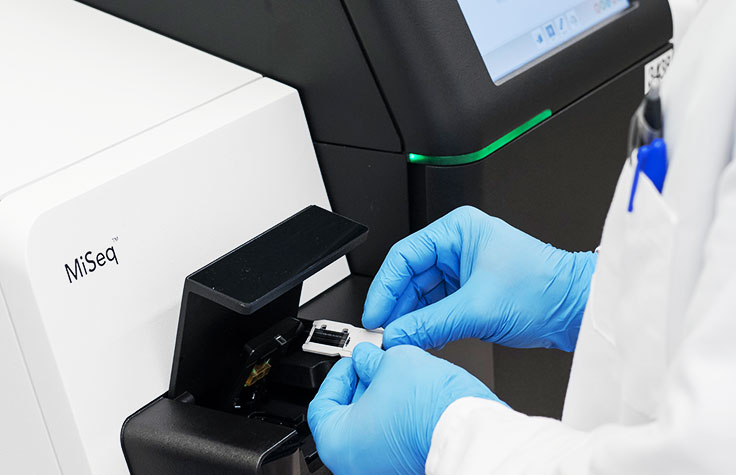

As a hypothesis-free method, NGS can distinguish between infectious disease strains that differ by as little as one SNP, and replace multiple tests.
Learn MoreRare Disease Genomics

See how genomics is driving a fundamental shift in rare disease research and analysis.
Learn MoreGene Sequencing with Targeted Panels
Learn how to sequence genes of interest to high depth with predesigned or custom panels.
Learn MoreSNP and SNV Genotyping
Compare techniques for detecting single nucleotide polymorphisms and variants to determine which approach is best for your needs. Learn how scientists are accelerating the ability to link variants to function and gain valuable insights into disease biology.
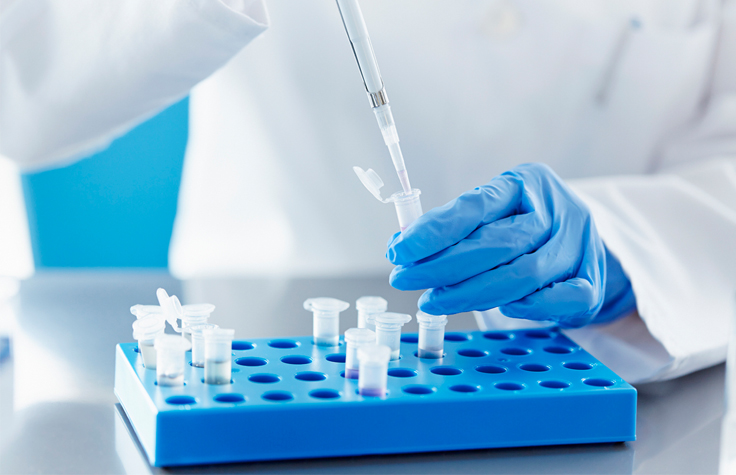

Additional Resources

Illumina DNA Prep Technology
Fast, integrated workflows for a wide range of applications.
Library Prep and Array Kit Selector
Find the right sequencing library preparation kit or microarray for your needs.

Customized panel content to fit your study needs

Germline CNV Analysis